Genstat Markdown
Reproducible Research with Genstat
2025-12-28
Reproducible research and Markdown
Reproducible research ensures that scientific findings can be independently verified by others using the same data and methods.
Markdown is a lightweight markup language used to format plain text into styled documents.
Markdown plays a key role in supporting reproducibility by allowing researchers to combine code, results, and narrative text in a clear, lightweight format.
Tools like R Markdown or Jupyter Notebooks use Markdown to embed executable code alongside explanations and outputs, making it easy to document workflows, share analyses, and regenerate results consistently across different environments.
Neither Simon nor I know much about Jupyter Notebooks, but we do know about R Markdown…
Why Genstat?

Some of you may know that this happened last November…
Why Genstat?
- Genstat has a long history in biometrics.
- It has strengths that R does not.
- It still has a large user base who may find this functionality useful.
Essential prerequistes
- (Access to) a valid Genstat licence.
- Genstat Messenger
- R and R Studio
- knitr, rmarkdown packages
Registering a custom language engine for knitr
- Yihui Xie, author of knitr among other things, provides documentation on how to create a custom engine so that knitr can render the input and output of a custom language in a nicely formatted document.
- It is also, perhaps, rather stereotypical in that it is a minimal example that covers the basics and tells you none of the complexities.
- Complexities include:
- How do I communicate with the other software?
- How do I keep a session open with the other software?
- How do I handle graphics?
- How do I handle default chunk options?
- How do I handle custom chunk options?
- …
Communication and session persistence
Users of R Markdown rely on a couple of features:
- The R interpreter/server is listening to their commands.
- The R session remembers previous commands.
This may seem obvious, but a naïve mechanism to pass code and results back and forth is batch processing. This process:
- Starts.
- Reads commands from a script.
- Writes results to file.
The script might just be a character string, and so might the outputs.
The session in this process has no persistence.
What do you mean “session has no persistence”?
It is very common to do something like this in R Markdown:
and then later you might embed this in the text:
This will not work unless the R session (or in this case the environment) containing the variable fit persists between the two chunks.
Genstat Messenger I
- Most users interact with Genstat through the GUI, which talks to the Genstat Server.
- It is possible to run Genstat in batch mode, but we lose session persistence.
- Enter Genstat Messenger.
Genstat Messenger Heroes


Genstat Messenger II
Genstat Messenger:
- Keeps a connection to the Genstat Server open.
- Relays commands to the Genstat Server.
- Receives output from the Genstat Server:
- Text output is transmitted as either plain text or HTML.
- Graphs are returned via Base64 encoding.
Communication architecture
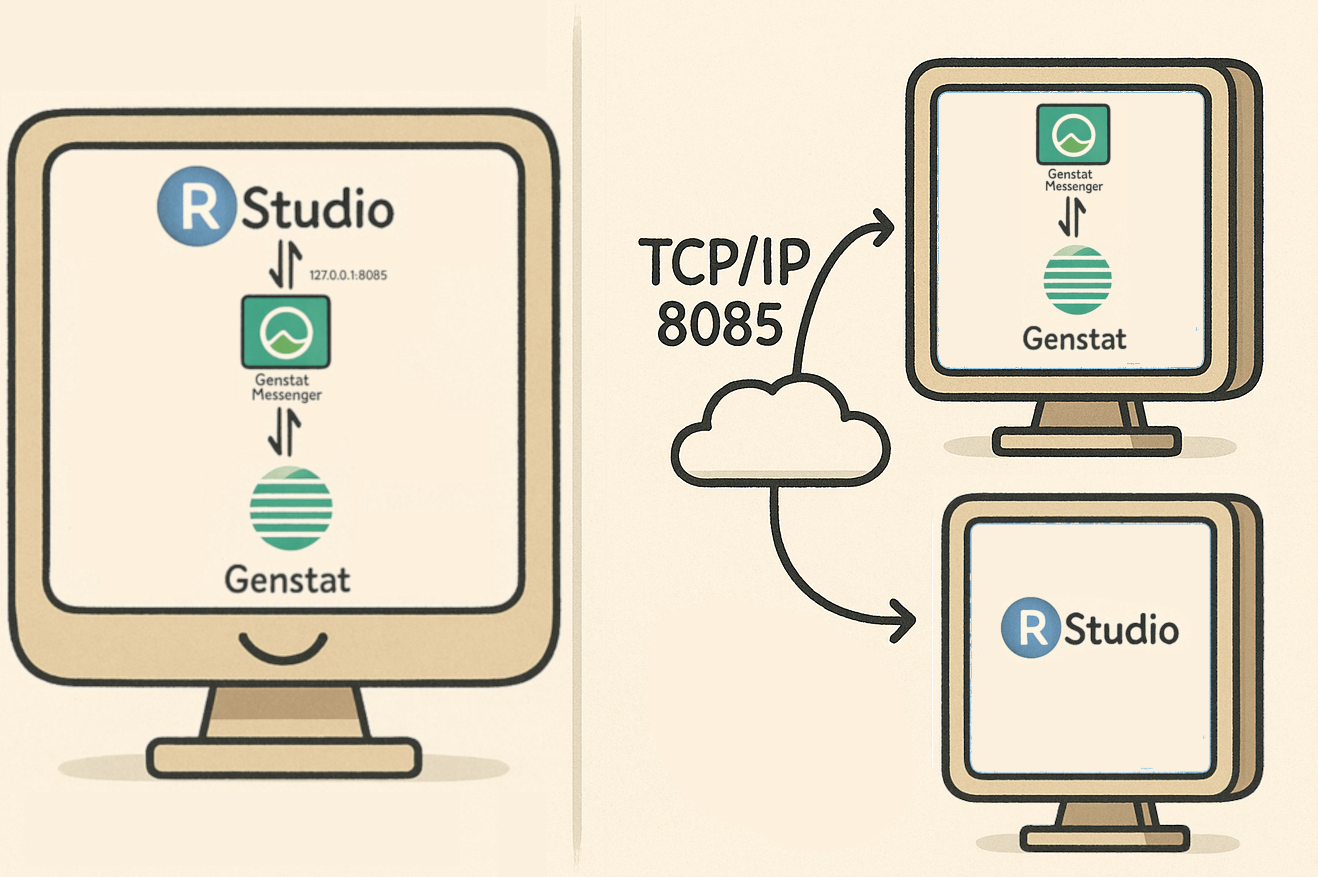
Okay, stop waffling. How do I use it?
Start Genstat Messenger
Open a new R Markdown (or Quarto) document in RStudio, and
set theknitr_enginefor Genstat:
Note: The choice of gs is arbitrary.
It is just a label we will use to tell knitr to use the Genstat engine.
- Write a Genstat chunk:
- Compile!
Does it work?
Yes, but that is not very exciting James! Let us take it up a notch:
What else have you got?
As an example we will analyse the built-in Wheat Trials data set. It contains information from a crop trial in New Zealand with 13 different cultivars.
IMPORT [PRINT=*] '%DATA%/WheatTrials.xlsx'; SHEET='Trial A1'
TREATMENTS Cultivar
BLOCKS Rep
ANOVA [PRINT=aov; FPROB=yes; PSE=LSD] Mean_Population_m2| Source of variation | d.f. | s.s. | m.s. | v.r. | F pr. |
| Rep stratum | 3 | 344.2 | 114.7 | 0.40 | |
| Rep.Units stratum | |||||
| Cultivar | 12 | 4354.8 | 362.9 | 1.26 | 0.283 |
| Residual | 36 | 10368.3 | 288.0 | ||
| Total | 51 | 15067.3 | |||
What about graphics?
Genstat users are likely to want to include graphics in their reports. We can do that too.
Calls to Genstat’s graphing procedures result in a PNG image, which are be displayed directly in the output document. These images are saved locally with filenames like ‘graph_1.png’, ‘graph_2.png’, etc.
Example:

Got anything else?
Saving Genstat tables for further manipulation
A lot of Genstat output is in tabular format. Equally, it is a very common in reproducible research to want to include figures from tables. For example, a sum-of-squares value or an \(F\)-statistic. The current implementation in gsengine allows for three different modes of table retrieval. The package can:
- Automatically assign all tables to a list to a name like
gs_tables_<chunklabel>. - Assign a single table (if there is only one) or a list to a given name.
- Return the first \(k\) tables to a list of \(k\) names.
We implement this through a chunk option saveTables.
- Option one is accessed through
saveTables=TRUE. - Option two might be something like
saveTables="aovTbls". - Option three might be something like
saveTables=c("aovTbl", "meanTbl").
A little bit of magic
We will run this chunk with label="wheat_anova", saveTables=TRUE.
This means that the ANOVA table will be stored in a list called gs_tables_wheat_anova. The first element of this list is the ANOVA table, and it is pretty unusable.
'data.frame': 10 obs. of 6 variables:
$ X1: chr "Source of variation" "" "Rep stratum" "" ...
$ X2: chr "d.f." "" "3" "" ...
$ X3: chr "s.s." "" "344.2" "" ...
$ X4: chr "m.s." "" "114.7" "" ...
$ X5: chr "v.r." "" "0.40" "" ...
$ X6: chr "F pr." "" "" "" ...
- attr(*, "gen_head_major")= chr "Analysis of variance"
- attr(*, "gen_head_minor")= logi NA
- attr(*, "gen_table_id")= chr "gs-table-4"A little bit of magic
However, we can help…
Analysis of Variance Table
Df Sum Sq Mean Sq F value Pr(>F)
Rep stratum 3 344.2 114.7 0.40
Cultivar 12 4354.8 362.9 1.26 0.283
Residual 36 10368.3 288.0
Total 51 15067.3 - This is, in some sense quite unusual behaviour, in that R Markdown was not really built to facilate tranport of objects between languages.
- However, given that all output comes back from Genstat as tables, it does offer a way to access information for reports.
Some caveats
- Currently all of our work is with HTML output from Genstat.
- PDF production is handled through
pandoc—specifically converting HTML to LaTeX. - This works okay for a report, but not so well for Beamer.
- PDF production is handled through
- There has not been extensive testing.
- There are some bugs in the save table code I have to fix.
Acknowledgements
- David Baird and VSNi
- Simon Urbanek
- Yuxiao Wang for taking this project on
- ChatGPT
- Emi Tanaka
James Curran, Genstat Markdown 2025-12-28 